Coding the future: next-gen bioinformatic tools for nanopore data
-
Project Leader

Dr Mike Clark+61 3 9035 3669
Project Details
Oxford Nanopore long-read sequencing is a cutting-edge approach with a number of unique features, including the real time sequencing of DNA or native RNA without the read length limits of traditional sequencing technologies. In addition, nanopore sequencing has extreme portability (take the sequencer to the sample) and can also detect nucleic acid base-modifications. This combination of unique attributes allows the use of nanopore sequencing in many novel (as well as existing) applications. Our lab has developed several new tools and applications for nanopore sequencing but there is much more that could be done to enable gene expression analysis, data visualisation, single cell sequencing and more. The opportunity exists for a student interested in bioinformatics or computational biology to develop new applications for nanopore sequencing to help address previously intractable biological questions.
Collaborators
Matt Richie, Walter and Eliza Hall Institute, Aus
Heejung Shim, University of Melbourne, Aus
Christine Wells, University of Melbourne, Aus
Research Publications
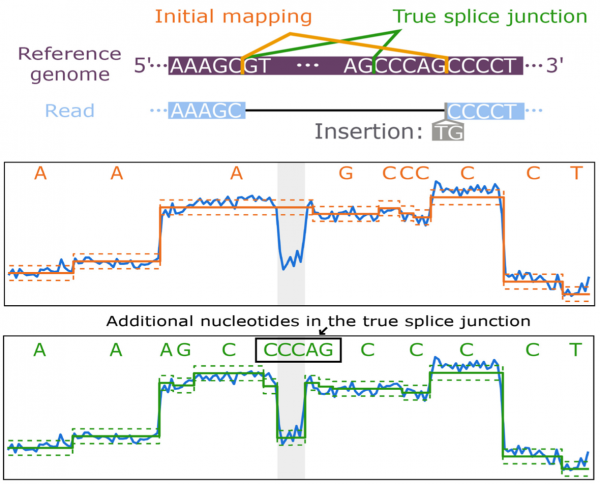
You, Y, Clark MB*, Shim H*.(2022). NanoSplicer: Accurate identification of splice junctions using Oxford Nanopore sequencing. Bioinformatics. btac359 (*Corresponding authors).
Gleeson J, Leger A, Prawer YDJ, Lane TA, Harrison PJ, Haerty W, Clark MB. (2022). Accurate expression quantification from nanopore direct RNA sequencing with NanoCount. Nucleic Acids Res. 50: e19
Research Group
Faculty Research Themes
School Research Themes
Cancer in Biomedicine, Biomedical Neuroscience, Stem Cells, Systems Biology, Molecular Mechanisms of Disease
Key Contact
For further information about this research, please contact the research group leader.
Department / Centre
MDHS Research library
Explore by researcher, school, project or topic.